Making AI Trustworthy: The importance of explainability in healthcare
The doctor's dilemma
A breath test shows elevated biomarkers, and the physician needs to explain the findings to their patient. "The AI detected an anomaly" does not provide much guidance either for the physician who needs to make clinical decisions or for the patient who wants to know what is wrong. In healthcare, diagnostic tools must deliver accurate predictions, while at the same time revealing the reasoning behind those predictions.
When a physician can see which specific biomarkers drove a result and understand why those markers matter, they can integrate AI findings with patient history, symptoms, and other diagnostic data. Without this transparency, even highly accurate AI remains difficult to trust and nearly impossible to validate.
The black box problem
Many modern AI systems can predict disease with impressive accuracy (1,2) yet they operate as black boxes: their input and output are known, but their reasoning remains elusive and difficult to explain in meaningful terms. This lack of transparency creates a fundamental trust problem in fields such as medicine and healthcare, where the stakes are particularly high. A model might correctly identify a disease with 95% confidence, but without understanding how it reaches its conclusions, physicians cannot verify its process, and patients cannot trust it with their health decisions.
For these reasons, regulatory bodies such as the FDA and medical device authorities in the EU increasingly demand interpretability before approving AI diagnostic tools. Physicians need to correlate AI findings with clinical presentation and patient history. Meanwhile, recent research in neural network interpretability (3,4) while promising, has not yet reached the level of transparency required for regulatory approval in medical devices.
The gap between what black box AI can do and what healthcare systems need remains substantial.
The breath analysis challenge
At RespiQ, each breath sample generates exceptionally high-dimensional data: information across hundreds of features in different domains, where each feature can reveal different chemical information. We are searching for disease biomarkers that exist at trace amounts, buried within overlapping features from hundreds of other volatile organic compounds.
This means every feature could represent a genuine biomarker or noise, and distinguishing between them demands rigorous scientific grounding. Some of these biomarkers are well-documented in existing literature; others are novel signals that our technology is uniquely positioned to uncover. In both cases, we need to trace each detected signal back to specific, interpretable molecular behaviours. When we identify a potential biomarker—whether known or previously unrecognised—we must pinpoint the precise features driving that detection and articulate why they are chemically meaningful. Without that interpretability, we have predictions without insight that cannot be clinically validated or trusted.
Our interpretability philosophy
RespiQ’s approach to AI development is built on a core principle: it should be possible to trace back every model decision to specific features, each with a clear scientific meaning. Our models are designed to identify features with real, measurable molecular meaning and not just to maximize prediction accuracy.
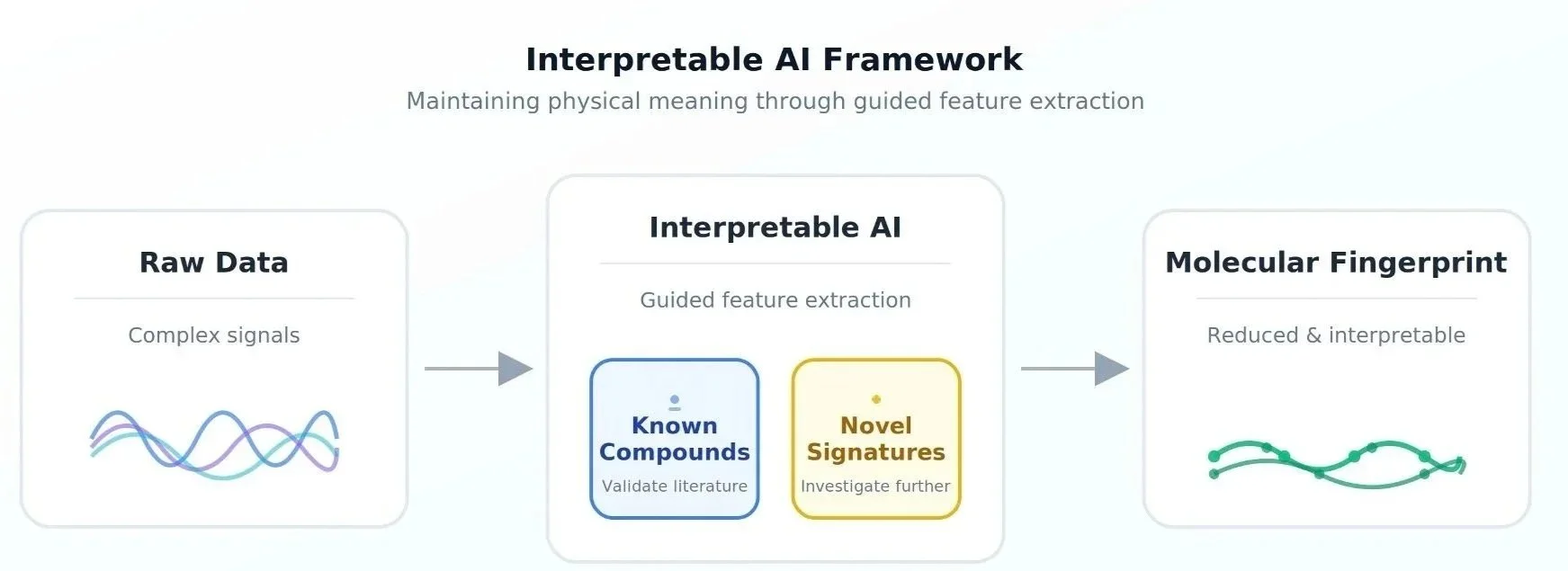
Our analytical framework serves a dual purpose: it makes the computational problem tractable while ensuring that the derived features maintain physical meaning. Rather than allowing the model to generate abstract mathematical representations, we guide it toward features that represent distinct patterns: unique molecular fingerprints that domain experts can validate. For known compounds, this means verifying that the AI identifies the same chemical markers already documented in scientific databases such as NIST or peer-reviewed literature, including our own published validation work (5,6). For novel biomarkers, it means the model reveals interpretable signatures that scientists can investigate, characterise, and ultimately explain through follow-up experimentation.
As we explained in our previous article on how AI powers breath analysis at RespiQ, the collaborative approach between experimental scientists and data scientists using interpretability as their common language enables a continuous feedback loop to not only apply existing knowledge but also actively contribute to scientific discovery while being transparent and auditable.
Feature extraction methodology that transforms complex signals into interpretable molecular fingerprints while preserving physical meaning through validation against known compounds and detection of novel signatures.
Why interpretability matters: from research to clinic
Consider how our device distinguishes between structurally similar volatile organic compounds—even when their signatures look almost identical —a challenge that defeats traditional analysis.
Our models identify specific features in the data and explain why those features are chemically meaningful. The reasoning chain is transparent: from raw sensor data we derive a molecular fingerprint informing a chemical interpretation, which in turn drives clinical insight. Whether the biomarker is well-established or newly discovered, clinicians can examine precisely which features drove the conclusion and assess the underlying logic.
This transparency is essential for scientific validation. It allows verification that the AI detects genuine molecular signals rather than spurious correlations, which is particularly important when the model identifies biomarkers that warrant further investigation. It also enables clinical adoption by allowing physicians to correlate AI findings with patient symptoms and history, seeing the chemical reasoning behind results rather than accepting an opaque verdict.
Looking ahead
RespiQ’s commitment to interpretable AI provides a strong scientific foundation as we expand our platform to detect additional compounds and disease signatures. This ensures that each new capability remains scientifically rigorous and generalizes effectively across diverse patient populations.
The same principles extend far beyond clinical diagnostics. In digestive health and nutrition monitoring, users can understand which specific fermentation byproducts their gut microbiome produces, enabling nuanced dietary choices rather than generic restrictions. In industrial applications—from critical logistics to chemical production—operators learn not just that something is wrong, but which molecular signatures indicate the problem, transforming alerts into actionable intelligence. Across all these domains, the principle is constant: understanding why matters as much as getting the right answer.
At RespiQ, we have built AI systems that explain their reasoning in interpretable, scientifically grounded terms, whether drawing on decades of established research or illuminating something entirely new. That approach earns trust from physicians, patients, regulators, and industrial users alike, and as our technology expands, interpretability remains the foundation ensuring that innovation and trust advance together.
RespiQ is developing AI-powered breath analysis technology for early disease detection and continuous health monitoring. Our interpretable AI approach combines advanced chemical sensing with explainable machine learning to achieve parts-per-billion sensitivity while maintaining transparency for clinical validation, regulatory approval, and physician adoption.
References
Ahsan, M. M., Luna, S. A. & Siddique, Z. Machine-Learning-Based Disease Diagnosis: A Comprehensive Review. Healthcare (Switzerland) 10, (2022). doi: https://doi.org/10.3390/healthcare10030541
Sadr, H. et al. Unveiling the potential of artificial intelligence in revolutionizing disease diagnosis and prediction: a comprehensive review of machine learning and deep learning approaches. Eur. J. Med. Res. 30, (2025). doi: https://doi.org/10.1186/s40001-025-02680-7
Zhang, Y., Tino, P., Leonardis, A. & Tang, K. A Survey on Neural Network Interpretability. IEEE Transactions on Emerging Topics in Computational Intelligence vol. 5 726–742 (2021). Preprint at https://doi.org/10.1109/TETCI.2021.3100641
Farah, L. et al. Assessment of Performance, Interpretability, and Explainability in Artificial Intelligence–Based Health Technologies: What Healthcare Stakeholders Need to Know. Mayo Clinic Proceedings: Digital Health vol. 1 120–138 (2023). Preprint at https://doi.org/10.1016/j.mcpdig.2023.02.004
Kramida, A. , R. Yu. , R. J. and N. A. T. NIST Atomic Spectra Database (version 5.12), [Online]. National Institute of Standards and Technology, Gaithersburg, MD (2024). doi: https://dx.doi.org/10.18434/T4W30F
van Poelgeest, J. et al. Exhaled volatile organic compounds associated with chronic obstructive pulmonary disease exacerbations—a systematic review and validation. J. Breath Res. 19, 026008 (2025). doi: 10.1088/1752-7163/adba06
Written by Chandan Sreedhara, 17/02/2026
Reviewed by Laura Verga, 24/02/2026